UCLA bioengineers and colleagues have created a new imaging system that advances dynamic imaging microscopy with artificial intelligence. The new system can reveal the details of biological processes in tiny tissue samples at a resolution of two thousandths of a millimeter and in slow motion at 200 frames per second.
A study outlining the advance was recently published in Nature Methods.
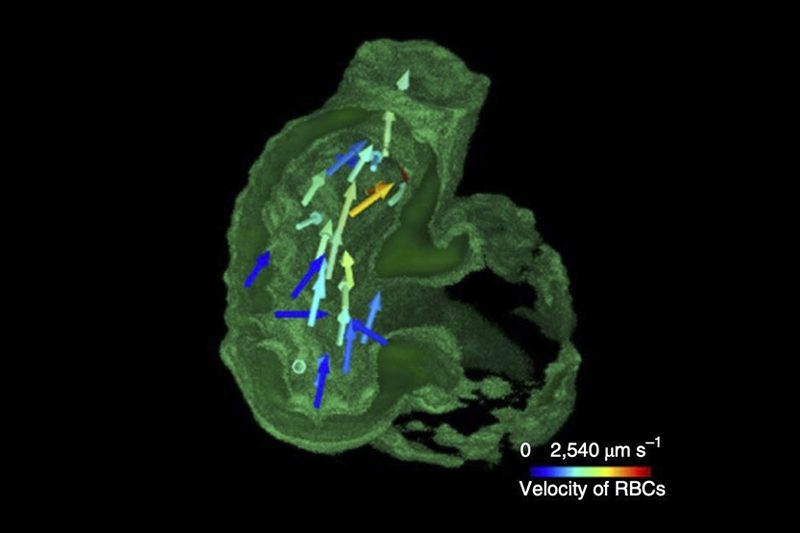
A recurring challenge in biology has been the extraction of spatiotemporal information from cell samples, as many millisecond-long, transient cellular processes occur in 3D tissues and across long time scales in space. Dynamics, such as flowing blood cells in the developing heart chambers, or rapidly moving neurons in the brain, are difficult to acquire as they require extremely high imaging speed, which remains an unmet optical challenge by existing microscopic techniques.
To tackle this problem, the researchers adopted a type of computational imaging tool, named light-field microscopy for 3D imaging, and enhanced it with a deep-learning neural network, which is a type of artificial intelligence-powered computing system modeled after how the human brain learns.
“This new system allows us to see biological events live in what is essentially five dimensions — the three dimensions of space, plus time and the molecular level dynamics as highlighted by color spectra,” said Dr. Tzung Hsiai, UCLA’s Maud Cady Guthman Professor of Cardiology. “For doctors and scientists, this could reveal the fine details of what’s happening in microscopic spaces and over millisecond-length time scales in a way that has never been done before. This advance can go a long way in helping find new insights to understand and treat diseases.”
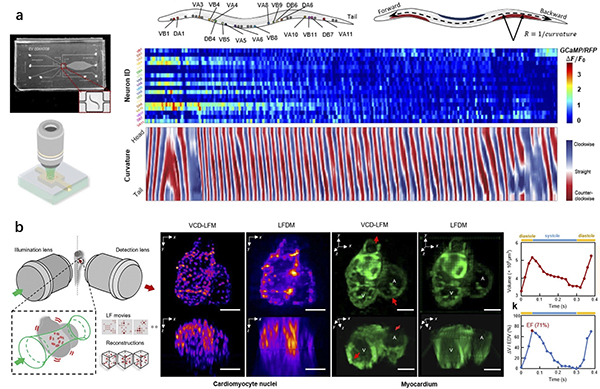
b. The illustration shows the heart muscles of fish embryos in action.
The new tool delivered a volumetric imaging at 200 cubic frames per second, revealing the transient processes inside a cell volume that measured 0.25 x 0.25 x 0.15 millimeters, or smaller than a grain of salt.
“Different from conventional microscopy, the tool reconstructed the 3D biological sample based on one snapshot through post-processing instead of scanning in the captured stage. The resulting temporal resolution of the images was drastically improved,” said lead author Zhaoqiang Wang, a doctoral student in bioengineering at the UCLA Samueli School of Engineering and member of Hsiai’s laboratory.
Compared to previously used light-field microscopy methods, Wang said the new technique also adopted a deep-learning technology to reconstruct images through a trained neural network model and thereby achieved better spatial resolution, image quality and processing throughput.
A neural network was first trained using 3D image stacks and corresponding light-field snapshots. The model was then used to infer the 3D reconstruction directly from the experimental light field, which records the dynamic process.
In a demonstration involving free-moving roundworms (C. elegans), the team used fluorescent tags to correlate neural signals with the roundworms’ motions. The researchers also recorded cardiac dynamics of embryonic zebrafish hearts, examining the flow of blood cells in synchrony with the contracting cardiomyocytes cells that make up the heart muscle.
The three co-corresponding authors on the study are Hsiai, Peng Fei and Shangbang Gao. Fei and Gao are both with the Huazhong University of Science and Technology in China. Hsiai, who directs the UCLA Cardiovascular Engineering Laboratory, holds faculty appointments in the Division of Cardiology and Department of Medicine at the UCLA David Geffen School of Medicine; the VA Greater Los Angeles Healthcare System; and in the Department of Bioengineering at UCLA Samueli.
Other authors on the study include Yichen Ding of UCLA Cardiovascular Engineering Laboratory; Lanxin Zhu, Hao Zhang, Guo Li, Chengqiang Yi, Yi Li and Yicong Yang of the Huazhong University; and Mei Zhen of the Mount Sinai Hospital at the University of Toronto in Canada.
The research was supported by the National Institutes of Health, the Department of Veterans Affairs and research agencies in China.